The pattern of rings that a tree lays down each year reflects how the tree has responded to its environment. So on the one hand, those rings tell us about variables such as disturbances and climate, and about age. On the other hand, underpinning those responses is the tree’s individual genome. The rings are one more aspect of the tree’s phenotype, its external appearance, and for most dendrochronologists (the scientists who study tree rings) the differences among individual specimens are just garbage, which they need to throw out somehow in order to detect the environmental signals common to all the individuals.
For GenTree, however, the garbage is gold. If we can relate an individual tree’s phenotype to its genotype, we can begin to understand what enables trees to evolve and adapt to changing environments, discovering which genes are most important and the effects they have. Having spent many hours measuring the tree rings of seven different species from 144 sites across Europe (see our photo story), the time had come to start the detailed analysis.
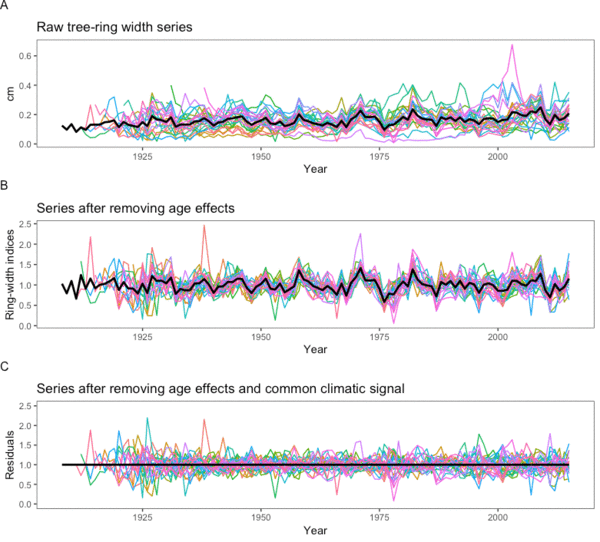
Geneticists and dendrochronologists from Philipps University in Marburg, Germany, and the Swiss Federal Research Institute WSL joined forces in two workshops at WSL to work out ways to use their collective knowledge to make the most of tree-ring data like that shown in Figure A. The key was to work out a shared strategy to remove the information common to all specimens (i.e., the climate effect on growth) in order to reveal the unique individual signal.
There are several steps. The first is common to conventional dendrochronology too, and consists of removing long-term changes that are mostly related to the age of the tree. This is called de-trending, and it gives us a new pattern for each tree. If you look closely at Figure B, which shows the data after de-trending, you can see that peaks are in the same place, but the relative size of some of them has changed. Having got rid of age effects, the main external influence on the growth patterns in Figure B is climate. Because climate has a similar effect for all individuals growing at a particular site, we can remove it by subtracting the average pattern, the black line in figure B, from each individual pattern.
This gives us the data in Figure C, where the average, the black line, is effectively the same every year. Now, the differences among the different individual patterns are the result of differences in the genetic make-up of the trees.
We have the gold we were looking for, but it will take a lot more work to realise its value. We will now test the procedure for different sites and species in order to validate the method, refine it and make sure we are getting the most possible information from the tree-ring phenotype. Once that is done, we will be able to move on and see how the genotype influences the phenotype, which in time will give us the tools to ensure that the management and use of forest genetic resources is sustainable and productive.